Phosphoproteomics on leftover material after RNA isolation
- Post by: OPL
- June 4, 2021
- Comments off
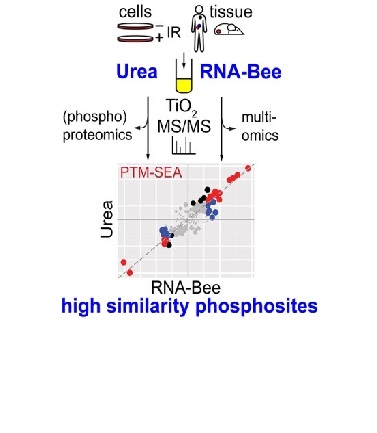
You throw away leftovers after RNA isolation? Save them and consider performing proteomics. The team of Connie Jimenez could show that RNA-Bee derived protein fractions are very suitable for phosphorylation analysis.
In daily practice, different types of biomolecules are usually extracted for large-scale “omics” analysis with tailored protocols. However, when sample material is limited, an all-in-one strategy is preferable. Although lysis of cells and tissues with urea is widely used for phosphoproteomic applications, DNA, RNA, and proteins can be simultaneously extracted from small samples using acid guanidinium thiocyanate–phenol–chloroform (AGPC). Use of AGPC for mass spectrometry–based phosphoproteomics was reported but has not yet been thoroughly evaluated against a classical phosphoproteomic protocol. We compared urea- with AGPC-based protein extraction, profiling phosphorylations in the DNA damage response pathway after ionizing irradiation of U2OS cells as proof of principle. On average we identified circa 9000 phosphosites per sample with both extraction methods. Moreover, we observed high similarity of phosphosite characteristics (e.g., 94% shared class 1 identifications) and deduced kinase activities (e.g., ATM, ATR, CHEK1/2, PRKDC). We furthermore extended our comparison to murine and human tissue samples yielding similar and highly correlated results for both extraction protocols. AGPC-based sample extraction can thus replace common cell lysates for phosphoproteomic workflows and may thus be an attractive way to obtain input material for multiple omics workflows, yielding several data types from a single sample.
For more information see: Feasibility of Phosphoproteomics on Leftover Samples After RNA Extraction With Guanidinium Thiocyanate
Funding for this work was provided by:
- NWO-Middelgroot project 91116017
- KWF project VU2013-6423
- German Research Foundation (DFG)
- European Union’s Horizon 2020 research and innovation program / Marie Skłodowska-Curie grant (agreement 722729)
